At a glance
Cancers with a high fatality rate include those typically diagnosed at a late stage, such as pancreatic cancer. Other cancers, such as breast cancer, are in fact a spectrum of unique subtypes driven by distinct events and characterized by unique biology. Some of these subtypes are more easily treated, while others are highly aggressive and refractory to standard treatments such as chemotherapy. Targeted therapies that specifically block the activity of the proteins driving the growth and survival of aggressive cancer subtypes can be more effective and safer than previous treatments. However, not all cancer drivers can be targeted through conventional drug development strategies. Some poor-prognosis cancers are also rare and understudied, meaning that we lack knowledge of how these tumours can be targeted. Large groups of patients therefore currently do not benefit from targeted therapies. Moreover, even the most advanced targeted therapies do not work equally well for all patients, with tumours often failing to respond as expected, or responding initially but then recurring or relapsing as resistant disease. Resistance to treatment is one of the most important factors currently limiting survivorship and reducing quality of life for cancer patients. It is often associated with metastasis, the process by which cancer cells leave their site of origin and spread to other organs. Notoriously difficult to treat, metastatic disease causes over 90% of cancer-associated deaths and is a major reason for the high fatality rate of cancers that are typically diagnosed late, when metastatic disease is already established.
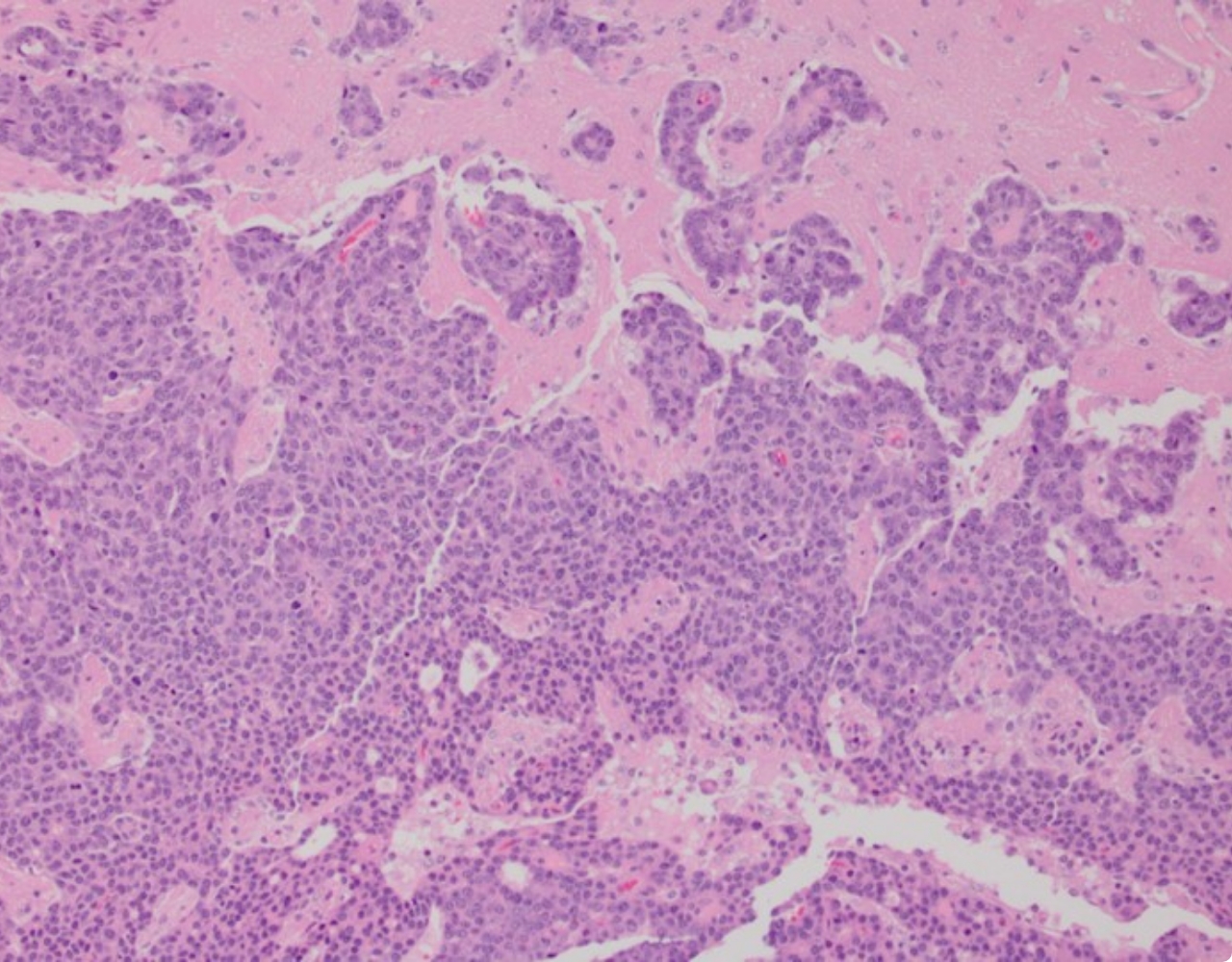
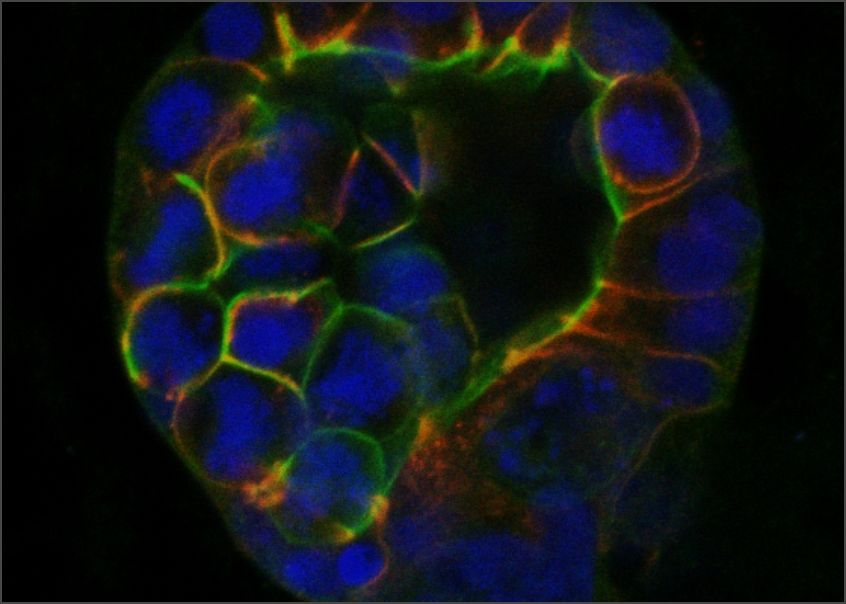
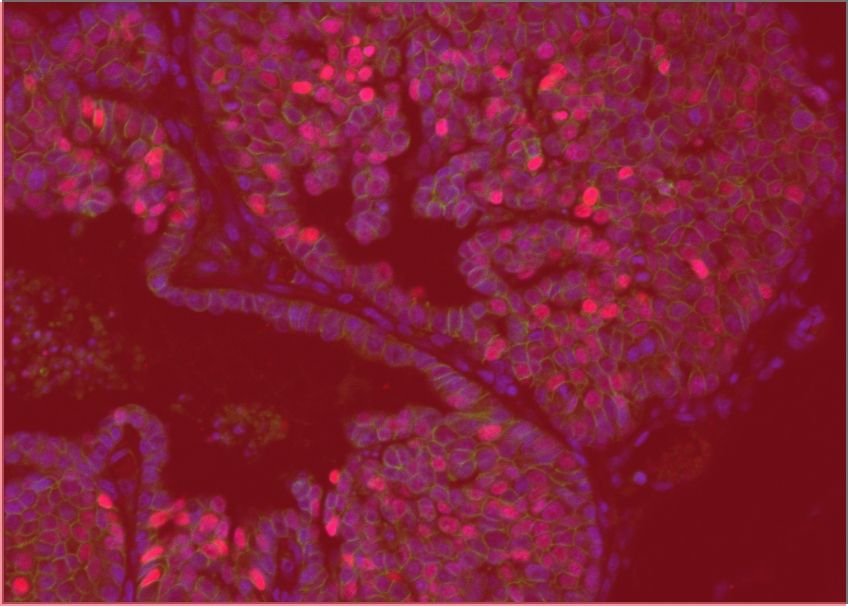
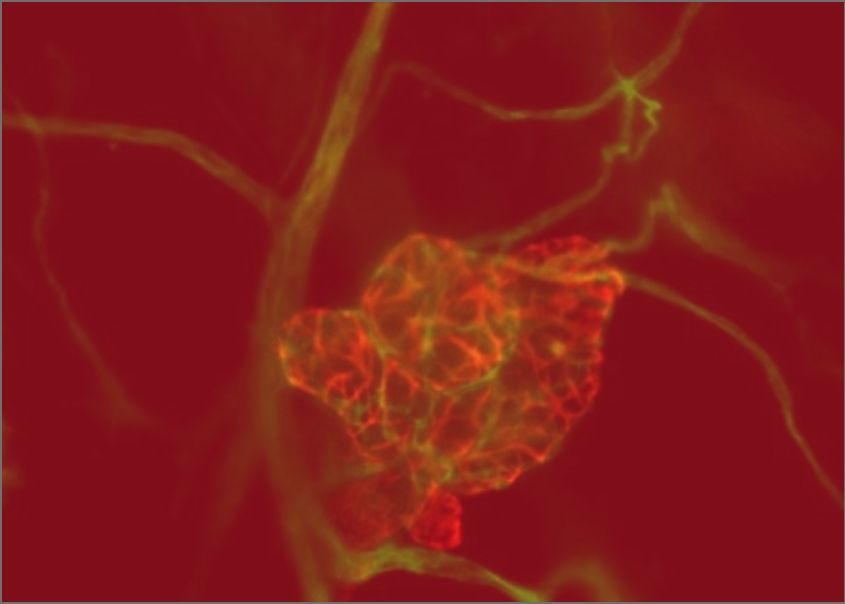
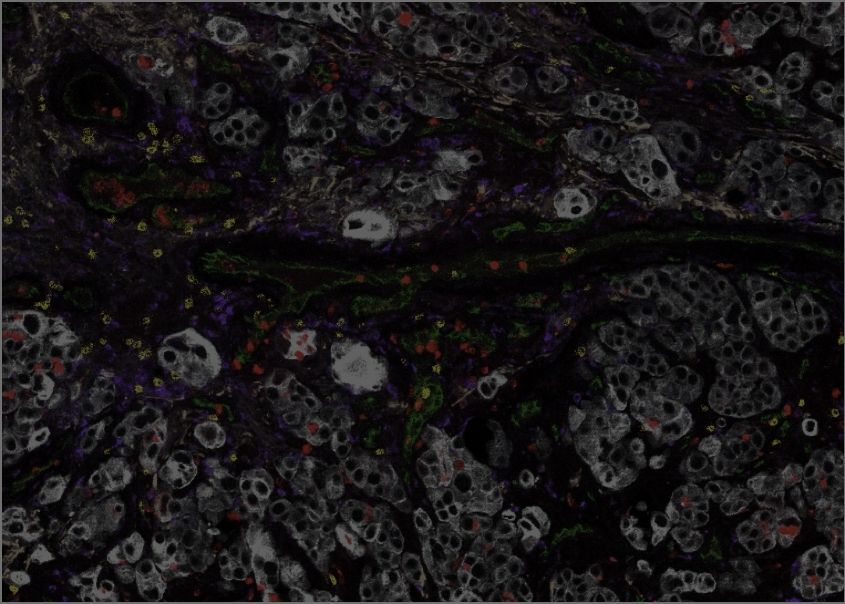
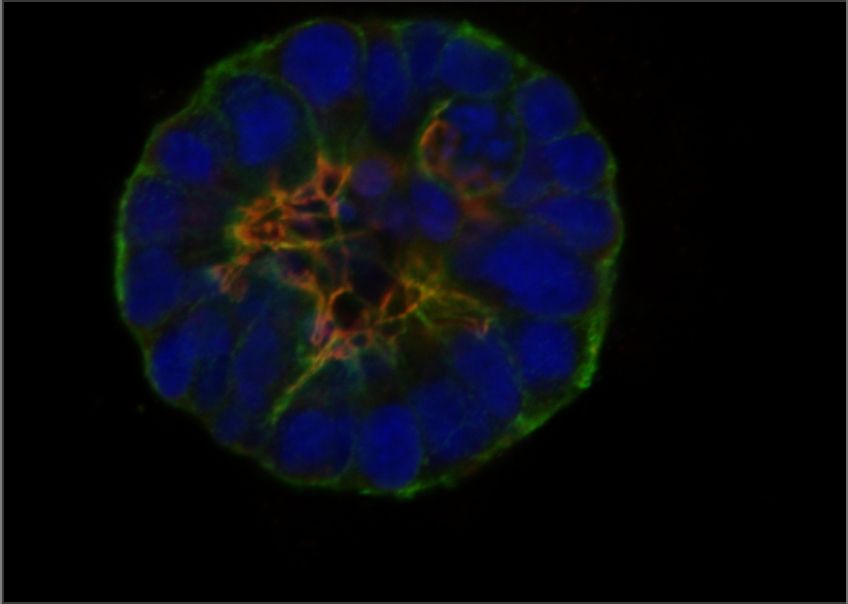
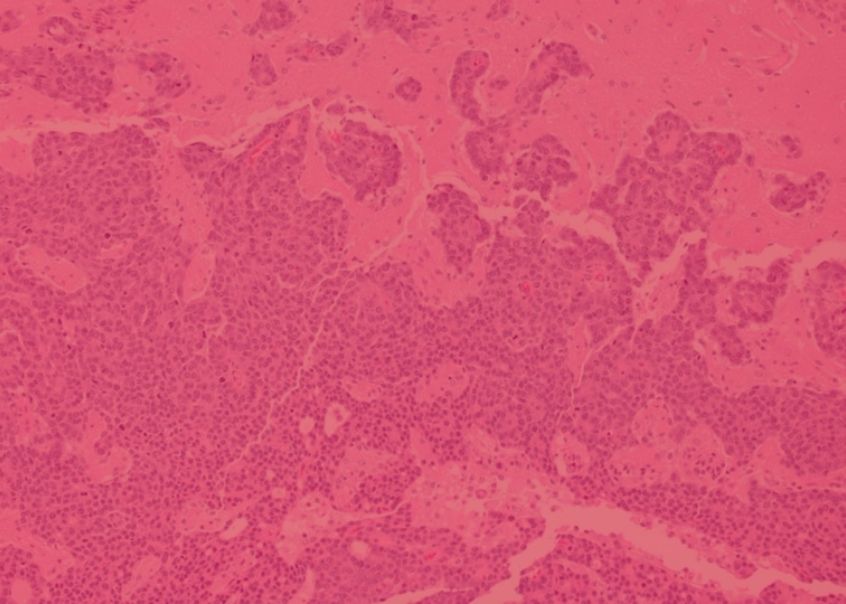
The GCI is a focal point of networks and consortia dedicated to precision medicine. By applying state-of-the-art technology and analytical methods to unique collections of clinical samples and data, we reveal the genomic, molecular and cellular features associated with metastatic progression and resistance to therapy. We have developed advanced techniques to recreate the features of high-fatality cancers in model organisms with a fully intact microenvironment, as well as technology to grow tumours in the laboratory directly from living samples surgically removed from patients. These highly accurate cancer models allow us to discover the mechanisms and vulnerabilities underlying resistant and metastatic disease and understand them at a fundamental level, leading to new and more effective therapeutic strategies for high-fatality cancers.
Areas of Focus
Team members
Our Discoveries
The “seed and soil” hypothesis is a long-standing idea explaining clinically observed patterns of metastasis, whereby specific cancer types metastasize preferentially to specific secondary organs. It posits that this occurs not by chance, but because each type of cancer cell is suited to survive and grow only in specific microenvironments. Work done at the GCI has provided unprecedented insight into this phenomenon by determining how cancer cells uniquely regulate their genetic and metabolic programs during metastasis to different organs. This has shed new light on the mechanisms of site-specific metastasis, identifying targetable vulnerabilities in the metastatic process at each site. Building on observations from clinical and pre-clinical studies, our investigators have also developed genetically engineered models that have uncovered how specific genes are involved in metastasis in the context of a fully intact microenvironment, including the immune system and all other stromal cell types. The findings of these studies have led to the development of therapeutic strategies, some of which have entered clinical trials, with the potential to transform outcomes for high-fatality cancers by blocking metastasis.
Osteoactivin promotes breast cancer metastasis to bone. Rose AA, et al. Mol Cancer Res. 2007 Oct;5(10):1001-14. doi: 10.1158/1541-7786.MCR-07-0119. PMID: 17951401
Phase I/II study of the antibody-drug conjugate glembatumumab vedotin in patients with locally advanced or metastatic breast cancer. Bendell J, et al., J Clin Oncol. 2014 Nov 10;32(32):3619-25. doi: 10.1200/JCO.2013.52.5683. Epub 2014 Sep 29. PMID: 25267761
Claudin-2 is selectively enriched in and promotes the formation of breast cancer liver metastases through engagement of integrin complexes. Tabariès S, et al., Oncogene. 2011 Mar 17;30(11):1318-28. doi: 10.1038/onc.2010.518. Epub 2010 Nov 15. PMID: 21076473
Claudin-2 promotes colorectal cancer liver metastasis and is a biomarker of the replacement type growth pattern. Tabariès S, et al., Commun Biol. 2021 Jun 2;4(1):657. doi: 10.1038/s42003-021-02189-9. PMID: 34079064
Identification of a Stat3-dependent transcription regulatory network involved in metastatic progression. Ranger JJ, et al., Cancer Res. 2009 Sep 1;69(17):6823-30. doi: 10.1158/0008-5472.CAN-09-1684. Epub 2009 Aug 18. PMID: 19690134
STAT3 Establishes an Immunosuppressive Microenvironment during the Early Stages of Breast Carcinogenesis to Promote Tumor Growth and Metastasis. Jones LM, et al., Cancer Res. 2016 Mar 15;76(6):1416-28. doi: 10.1158/0008-5472.CAN-15-2770. Epub 2015 Dec 30. PMID: 26719528
GCI investigators have built a world-leading research program on high-fatality breast cancers, including metastatic, drug resistant, and rare subtypes. We are pioneers in developing genetically engineered models of poor-prognosis subtypes known as HER2-positive and triple-negative breast cancer (TNBC). These models have been used at the GCI and around the world to understand how these breast cancers form and progress, leading to many important discoveries and the development of new therapies. We established the first biobank of breast cancer patient samples and data in Quebec, now part of the FRQS-supported Reseau du Recherche en Cancer (RRCancer), and a precision oncology research pipeline for breast cancer that links the beside and the bench. This unique resource allows patient samples to be developed into models maintained as living tumours in mice (patient-derived xenografts) or grown in the lab (patient-derived organoids), where we use sophisticated technology, such as 3D printing of cells and extracellular matrix components, to recreate the tumour microenvironment. Overall, combining our expertise in cancer modeling with genomics and associated technologies, computational biology and AI, innovative molecular and cell biology techniques, advanced drug screening and functional genomics — an approach that can systematically inactivate each gene in the entire genome — allows us to discover new therapeutic targets to prevent progression and overcome resistance in these aggressive breast cancer subtypes.
Met induces mammary tumors with diverse histologies and is associated with poor outcome and human basal breast cancer. Ponzo MG, Lesurf R, et al. Proc Natl Acad Sci U S A. 2009 Aug 4;106(31):12903-8. doi: 10.1073/pnas.0810402106. Epub 2009 Jul 17. PMID: 19617568
Met synergizes with p53 loss to induce mammary tumors that possess features of claudin-low breast cancer. Knight JF, Lesurf R, et al. Proc Natl Acad Sci U S A. 2013 Apr 2;110(14):E1301-10. doi: 10.1073/pnas.1210353110. Epub 2013 Mar 18. PMID: 23509284
Chemogenomic profiling of breast cancer patient-derived xenografts reveals targetable vulnerabilities for difficult-to-treat tumors. Savage P, et al. Commun Biol. 2020 Jun 16;3(1):310. doi: 10.1038/s42003-020-1042-x. PMID: 32546838
Reduction of Global H3K27me3 Enhances HER2/ErbB2 Targeted Therapy. Hirukawa A, et al. Cell Rep. 2019 Oct 8;29(2):249-257.e8. doi: 10.1016/j.celrep.2019.08.105. PMID: 31597089
Targeting EZH2 reactivates a breast cancer subtype-specific anti-metastatic transcriptional program. Hirukawa A, et al. Nat Commun. 2018 Jun 29;9(1):2547. doi: 10.1038/s41467-018-04864-8. PMID: 29959321
Folliculin impairs breast tumor growth by repressing TFE3-dependent induction of the Warburg effect and angiogenesis. El-Houjeiri L, Biondini M, et al. J Clin Invest. 2021 Nov 15;131(22):e144871. doi: 10.1172/JCI144871. PMID: 34779410
ERRα mediates metabolic adaptations driving lapatinib resistance in breast cancer. Deblois G, et al. Nat Commun. 2016 Jul 12;7:12156. doi: 10.1038/ncomms12156. PMID: 27402251
Metastatic melanoma is a highly aggressive and difficult-to-treat disease, the incidence of which is unfortunately increasing. Most melanomas are driven by genetic mutations affecting the activity of a specific cellular signaling pathway, the RAS-MAP kinase pathway, which is a master regulator of cell growth, division, and survival. Although precision medicine strategies targeting the RAS-MAP kinase pathway lead to responses in many patients, their effectiveness is inevitably limited by resistance. GCI investigators are explaining this heterogeneity by leading the world’s most comprehensive genomic studies of metastatic melanoma, using large numbers of patient samples to map the genomic landscape of this disease. As these studies reveal new molecular subtypes and genomic drivers of melanoma, our fundamental investigation of the underlying biology matches molecular subtypes with the most effected targeted therapeutic strategies.
A landscape of driver mutations in melanoma. Hodis E, Watson IR, et al. Cell. 2012 Jul 20;150(2):251-63. doi: 10.1016/j.cell.2012.06.024. PMID: 22817889
Genomic Classification of Cutaneous Melanoma. Cancer Genome Atlas Network. Cell. 2015 Jun 18;161(7):1681-96. doi: 10.1016/j.cell.2015.05.044. PMID: 2609104
The RAC1 P29S hotspot mutation in melanoma confers resistance to pharmacological inhibition of RAF. Watson IR, et al. Cancer Res. 2014 Sep 1;74(17):4845-4852. doi: 10.1158/0008-5472.CAN-14-1232-T. Epub 2014 Jul 23. PMID: 25056119
Dual MAPK Inhibition Is an Effective Therapeutic Strategy for a Subset of Class II BRAF Mutant Melanomas. Dankner M, et al. Clin Cancer Res. 2018 Dec 15;24(24):6483-6494. doi: 10.1158/1078-0432.CCR-17-3384. Epub 2018 Jun 14. PMID: 29903896
Multi-omic analysis reveals significantly mutated genes and DDX3X as a sex-specific tumor suppressor in cutaneous melanoma. Alkallas R, et al. Nat Cancer. 2020 Jun;1(6):635-652. doi: 10.1038/s43018-020-0077-8. Epub 2020 Jun 22.PMID: 35121978
Melanomas with concurrent BRAF non-p.V600 and NF1 loss-of-function mutations are targetable by BRAF/MEK inhibitor combination therapy. Rajkumar S, et al. Cell Rep. 2022 Apr 5;39(1):110634. doi: 10.1016/j.celrep.2022.110634. PMID: 35385748
Pancreatic cancer is one of the most aggressive tumour types, lacking effective therapies and typically diagnosed at a late stage when surgery is no longer a curative option. Research at the GCI, in partnership with the McGill University Health Centre (MUHC) and the Marathon of Hope Cancer Centres Network (MOHCCN) of the Terry Fox Research Institute (TFRI), includes innovative genomics, biobanking and patient-derived modeling programs that are revealing the drivers of this deadly disease and uncovering the genetics underlying inherited risk of pancreatic cancer. Our investigators have developed the Quebec Pancreas Cancer Study (QPCS), a clinical and translational pancreatic cancer research resource encompassing over 1000 families with a history of pancreatic cancer, including a knowledge and biospecimen bank with a cutting-edge patient-derived modeling platform. Through the QPCS and leadership within high-impact multi-centre Canadian and international initiatives and consortia including the Pancreatic Cancer Genetic Epidemiology Consortium (PACGENE), the Pancreatic Cancer Early Detection Consortium (PRECEDE), the Pancreatic Canadian Oncology Network (PancOne), and Enhanced Pancreatic Cancer Profiling for Individualized Care (EPPIC), a TFRI Translational Research Program, we are enabling new precision medicine strategies for pancreatic cancer through biomarker discovery and implementation, cutting-edge pre-clinical studies, and clinical trials that are making a difference for patients.
Oncology clinic-based germline genetic testing for exocrine pancreatic cancer enables timely return of results and unveils low uptake of cascade testing. Wang Y, et al. J Med Genet. 2022 Aug;59(8):793-800. doi: 10.1136/jmedgenet-2021-108054. Epub 2021 Sep 23. PMID: 34556502
A Preclinical Trial and Molecularly Annotated Patient Cohort Identify Predictive Biomarkers in Homologous Recombination-deficient Pancreatic Cancer. Wang Y, et al. Clin Cancer Res. 2020 Oct 15;26(20):5462-5476. doi: 10.1158/1078-0432.CCR-20-1439. Epub 2020 Aug 14
A region-based gene association study combined with a leave-one-out sensitivity analysis identifies SMG1 as a pancreatic cancer susceptibility gene. Wong C, et al. PLoS Genet. 2019 Aug 30;15(8):e1008344. doi: 10.1371/journal.pgen.1008344. eCollection 2019 Aug. PMID: 31469826
Establishing a clinic-based pancreatic cancer and periampullary tumour research registry in Quebec. Smith AL, et al. Curr Oncol. 2015 Apr;22(2):113-21. doi: 10.3747/co.22.2300. PMID: 25908910
Our ability to make an impact on research into rare cancer subtypes and discover new treatments is illustrated by a dynamic program on small cell carcinoma of the ovary, hypercalcemic type (SCCOHT), a rare form of ovarian cancer that is typically diagnosed in women under 40 years old, with long-term survival of only 30%. Using our patient-derived cancer modeling platform, in collaboration with McGill’s Lady Davis Institute and investigators from Greece, we derived a SCCOHT model directly from a sample surgically removed from a young patient. Applying our world-leading expertise in the functional characterization of cancer genomes, we used this model to discover a therapeutic target with a clinically approved targeted therapy, establishing a new strategy that could move rapidly into the clinic. Further studies revealed that other cancers with a genetic profile like SCCOHT, including some lung cancers, responded to the same treatment, leading to a nation-wide clinical trial in patients suffering from these cancers.
CDK4/6 inhibitors target SMARCA4-determined cyclin D1 deficiency in hypercalcemic small cell carcinoma of the ovary. Xue Y, et al. Nat Commun. 2019 Feb 4;10(1):558. doi: 10.1038/s41467-018-06958-9. PMID: 30718512
Small-Cell Carcinoma of the Ovary, Hypercalcemic Type-Genetics, New Treatment Targets, and Current Management Guidelines. Tischkowitz M, Huang S, et al. Clin Cancer Res. 2020 Aug 1;26(15):3908-3917. doi: 10.1158/1078-0432.CCR-19-3797. Epub 2020 Mar 10. PMID: 32156746
SMARCA4 loss is synthetic lethal with CDK4/6 inhibition in non-small cell lung cancer. Xue Y, et al. Nat Commun. 2019 Feb 4;10(1):557. doi: 10.1038/s41467-019-08380-1. PMID: 30718506
SMARCA4/2 loss inhibits chemotherapy-induced apoptosis by restricting IP3R3-mediated Ca2+ flux to mitochondria. Xue Y, et al. Nat Commun. 2021 Sep 13;12(1):5404. doi: 10.1038/s41467-021-25260-9. PMID: 34518526